LIMMS-UGSF-CANTHER-IEMN project
Researchers from Lille and Tokyo have joined forces to design specific sensors for proteins in the SARS-CoV-2 virus.
Simulations aim to elucidate the molecular mechanisms of recognition and interaction of these proteins.
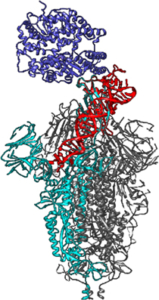
In a project involving experimental and modelling teams from LIMMS (Tokyo), IEMN, UGSF and CANTHER (Lille), we are using large-scale molecular simulations to design nanometric quantum sensors, aimed at recognising the 'spike' protein and the N protein of the SARS-CoV-2 virus, by means of ssDNA aptamers. Very cheap, easy to produce in large quantities, extremely biocompatible and with virtually no undesirable effects, aptamers are able to bind specifically to targets and modulate or block their activity, just like natural antibodies. Generated by an in vitro selection process from pools of random sequence oligonucleotides (the SELEX technique), a typical aptamer is 10 to 30 kDa in size (around 30 to 60 nucleotides of DNA or RNA), binds its target with sub-nanomolar affinity and can discriminate between closely related targets. Our molecular simulations elucidate the microscopic details of how DNA interacts with key viral proteins. In addition, in the context of innovative CoVid-19 therapies, we are investigating the possibility that the same DNA aptamers could actively block, or at least slow down, the opening mechanism of protein S, thereby suppressing or greatly reducing the cell's binding to receptors, which is the main means of transmission of infection by the virus.
.
This work is supported by the GENCI national supercomputer initiative.